![PDF] Clinotator: analyzing ClinVar variation reports to prioritize reclassification efforts | Semantic Scholar PDF] Clinotator: analyzing ClinVar variation reports to prioritize reclassification efforts | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/a7c0e187c02501c23bc0ac6fa66512fd0cb1c655/4-Table1-1.png)
PDF] Clinotator: analyzing ClinVar variation reports to prioritize reclassification efforts | Semantic Scholar
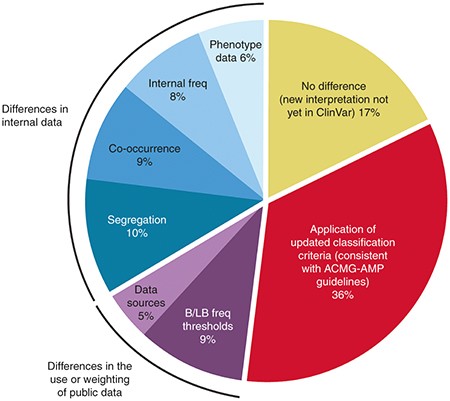
Clinical laboratories collaborate to resolve differences in variant interpretations submitted to ClinVar | Genetics in Medicine
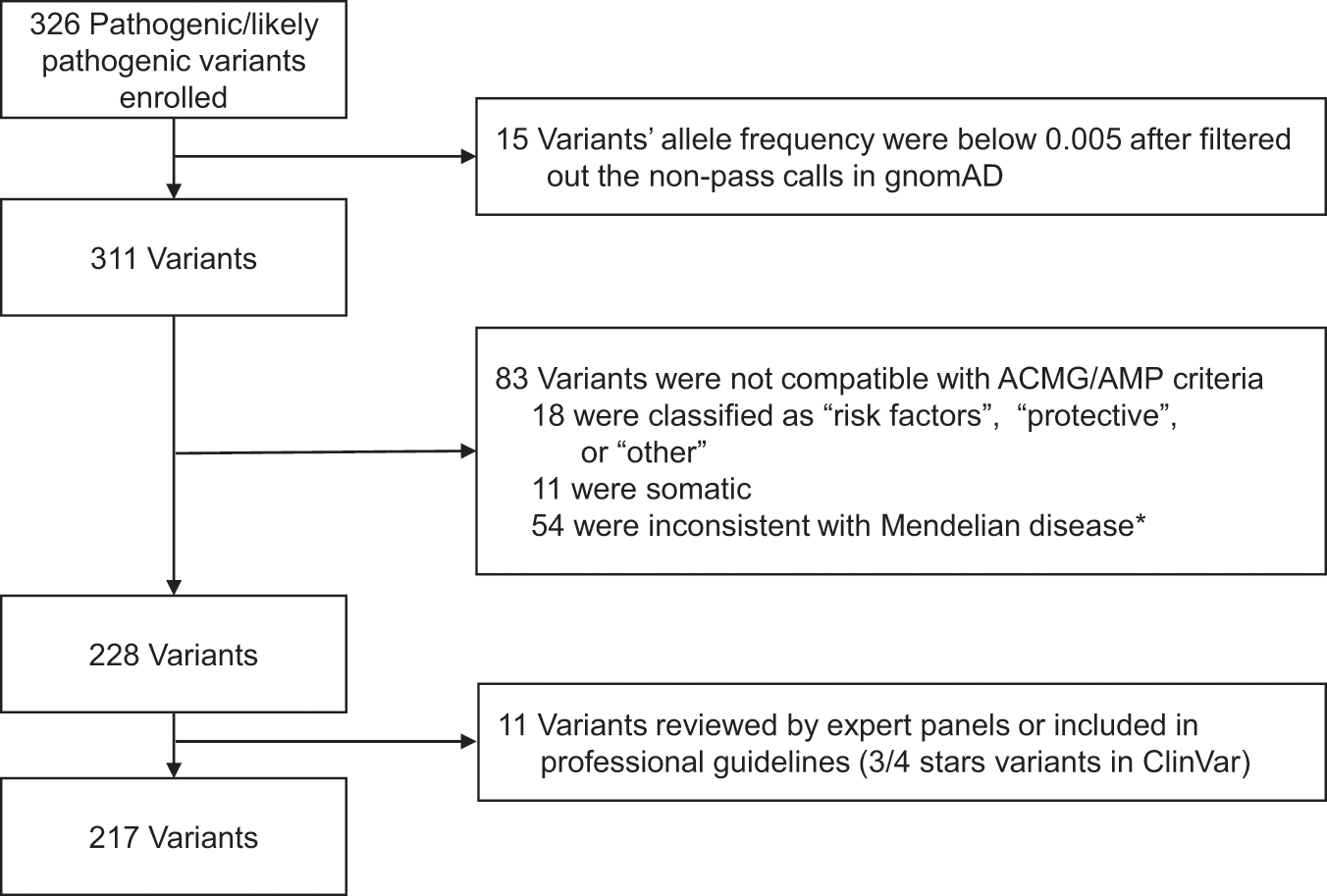

Reinterpretation of common pathogenic variants in ClinVar revealed a high proportion of downgrades | Scientific Reports

Assessment of an automated approach for variant interpretation in screening for monogenic disorders: A single‐center study - Gall - Molecular Genetics & Genomic Medicine - Wiley Online Library

CIViC is a community knowledgebase for expert crowdsourcing the clinical interpretation of variants in cancer | Nature Genetics
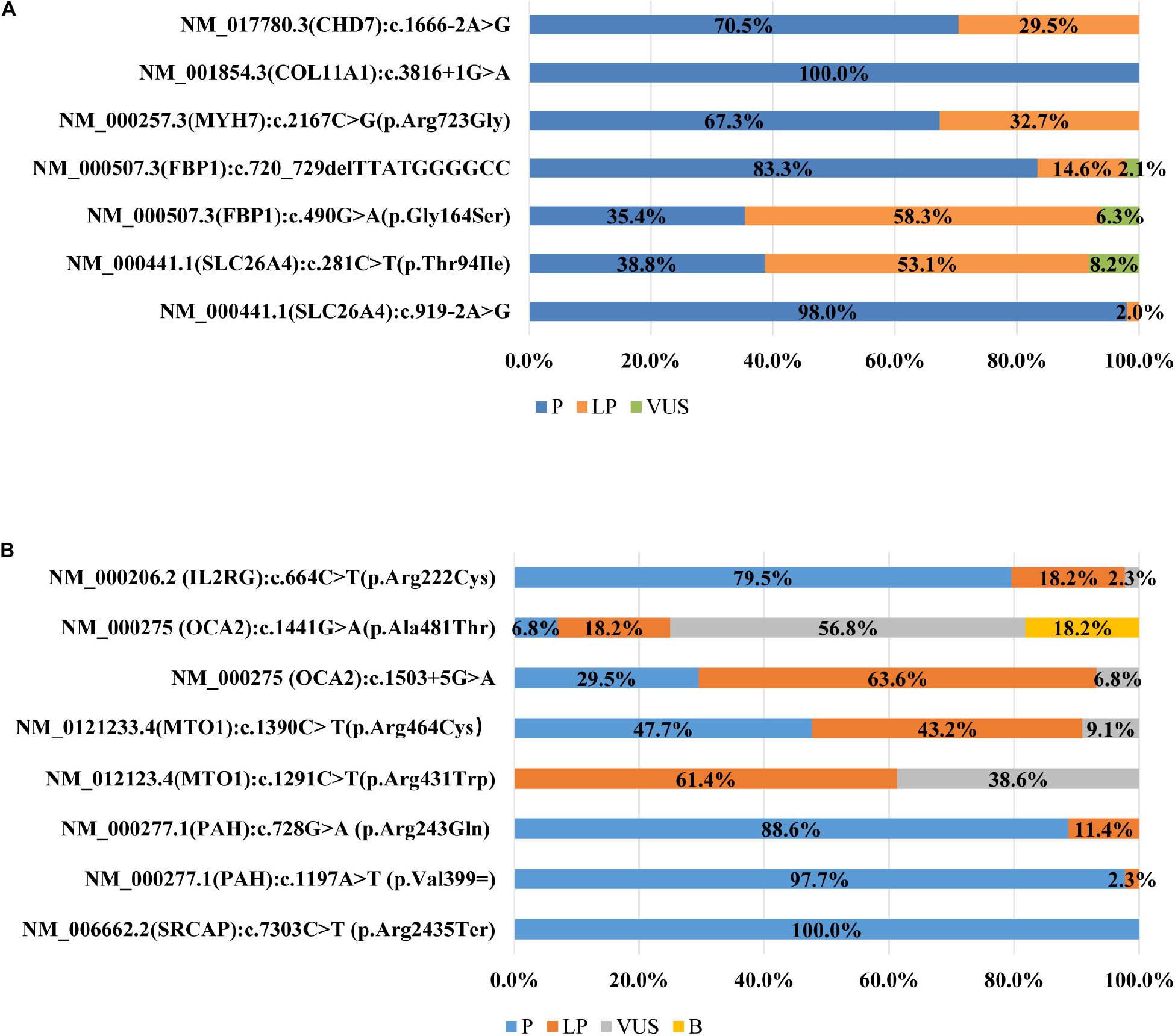
Frontiers | An Initial Survey of the Performances of Exome Variant Analysis and Clinical Reporting Among Diagnostic Laboratories in China
![PDF] ClinVar: public archive of relationships among sequence variation and human phenotype | Semantic Scholar PDF] ClinVar: public archive of relationships among sequence variation and human phenotype | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/ed95339eb652c67570cd2b0093a2758b8139e9e1/4-Figure1-1.png)
PDF] ClinVar: public archive of relationships among sequence variation and human phenotype | Semantic Scholar
![PDF] Simple ClinVar: an interactive web server to explore and retrieve gene and disease variants aggregated in ClinVar database | Semantic Scholar PDF] Simple ClinVar: an interactive web server to explore and retrieve gene and disease variants aggregated in ClinVar database | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/253417a4bf8f7b320ab286223e372e99283d6dd4/3-Figure1-1.png)
PDF] Simple ClinVar: an interactive web server to explore and retrieve gene and disease variants aggregated in ClinVar database | Semantic Scholar
Refinement of the clinical variant interpretation framework by statistical evidence and machine learning

Cells | Free Full-Text | A Next Generation Sequencing-Based Protocol for Screening of Variants of Concern in Autism Spectrum Disorder
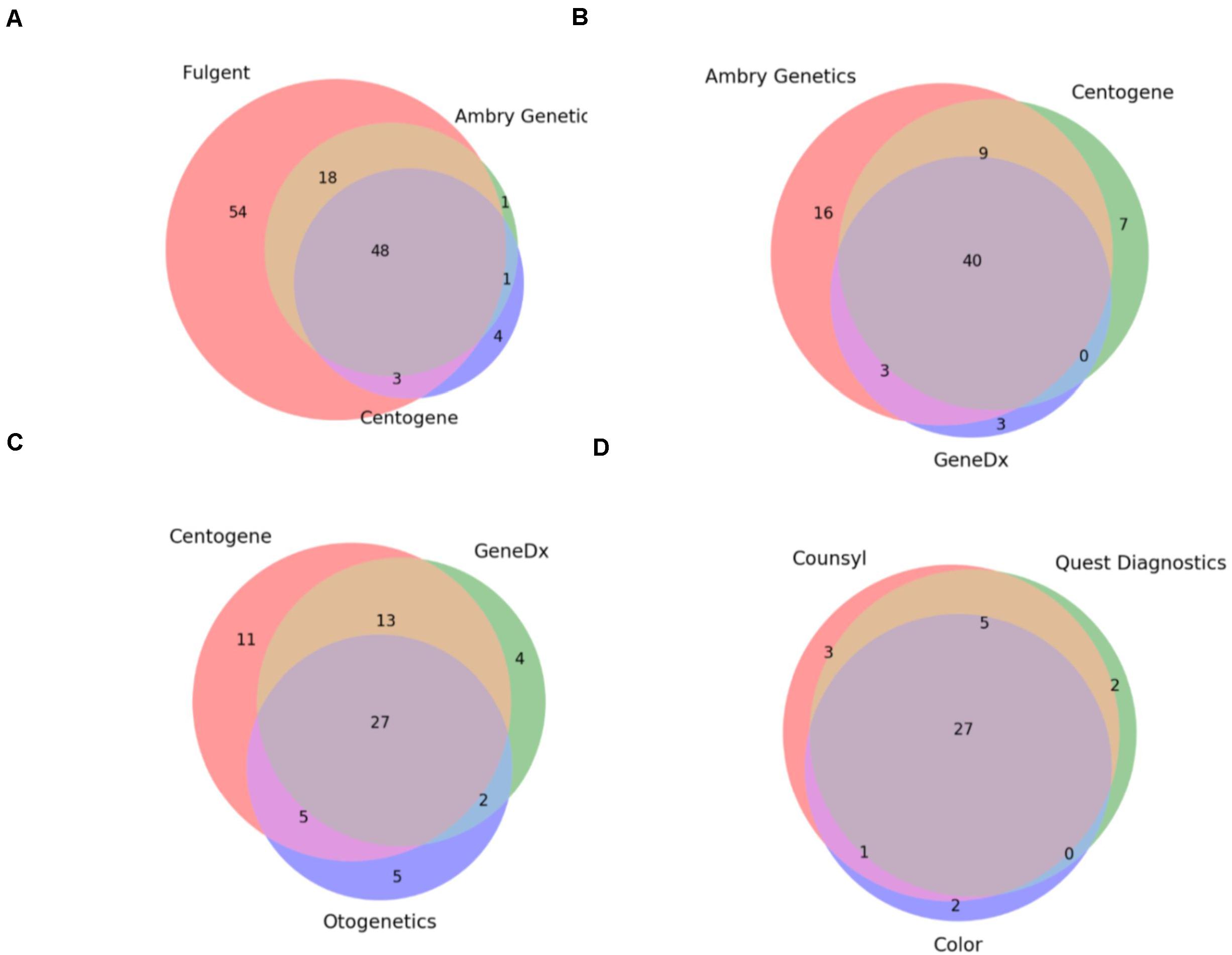
Frontiers | Mastermind: A Comprehensive Genomic Association Search Engine for Empirical Evidence Curation and Genetic Variant Interpretation
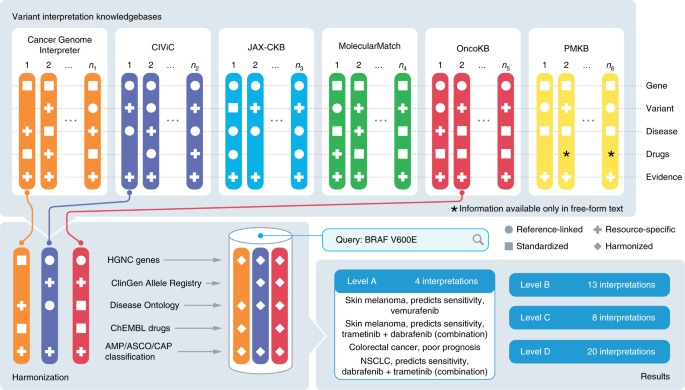







![PDF] ClinVar: public archive of interpretations of clinically relevant variants | Semantic Scholar PDF] ClinVar: public archive of interpretations of clinically relevant variants | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/a77e1886e8d8659f89c6dfacbbf339cb3c5c0672/4-Figure2-1.png)